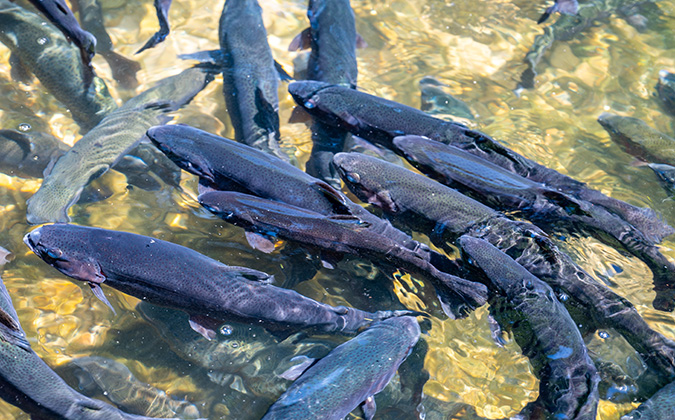
Screening work sheds new light on disease risks in northeast Pacific salmon farming
The use of cutting-edge technology has helped Canadian researchers carry out some of the most detailed screenings to date of pathogens affecting Atlantic salmon aquaculture.
Their multi-year research project, sampling four salmon farms in British Columbia, used high-throughput qPCR, a genetic profiling technology, to screen for 58 infectious agents.
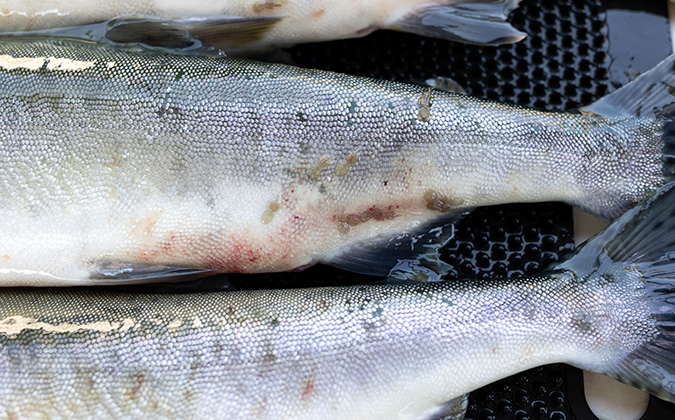
Among a range of findings, the scientists found Tenacibaculum maritimum, the causative agent of tenacibaculosis, prevalent among dead and dying fish. Two viruses only recently observed in aquaculture in the region, Atlantic salmon calicivirus (ASCV) and cutthroat trout virus-2 (CTV-2), were also detected.
Infection patterns detailed
Levels of infectious agents observed were highest among smolts coming out of freshwater hatcheries, the researchers explained.
“While the effect dissipated once fish entered the marine environment, it is clear that hatchery fish are dying with — or of — elevated levels of infection. The patterns we observed likely reflect the transition from freshwater to saltwater, with a coincident shift in infective-agent communities,” they said.
“Smoltification has also been associated with immune depression, and elevated infectious burden around the time of ocean entry may reflect this.”
Some unusual patterns were observed in other disease-causing agents. The scientists observed two marine pathogens (Kudoa thyrsites and T. maritimum) in freshwater settings, and one freshwater agent (Flavobacterium psychrophilum) in the ocean.
The unexpected marine species may have been due to hatcheries introducing saltwater in the weeks before sea transfer, they suggested, while in the case of the F. psychrophilum, it was not the first time it has been observed in a marine setting, and its survival has been proven in brackish water.
Salmon aquaculture’s complex picture
The size of the study brought considerable new data, but also set the groundwork for future work, with a number of observations requiring further explanation.
“The data and analyses we have presented provide a unique look into the epidemiology of farmed salmon populations, and wildlife/livestock diseases generally. No past studies have had access to multiple farmed-salmon cohorts, throughout their production cycles, with the capacity to molecularly screen for a large suite of infectious agents,” they explained.
As well as highlighting the presence of individual pathogens, the study demonstrated signs of coinfection that “clearly differed from random chance,” the scientists said. However, they were cautious in making wider inferences from the work, which they stressed was correlative.
“Not all infective agents cause disease, and even agents that do can be present long before—or long after—clinical symptoms. Our work presents only a piece of the puzzle in what is a multifaceted, complex scenario of shared wildlife/livestock disease in salmon aquaculture,” they said.
Screening helps preempt disease outbreaks
The researchers stated a case for the importance of studying pathogens in an aquaculture setting, suggesting that understanding low-intensity infections, and not just disease outbreaks, helps better manage disease in general.
Such studies also help understand how diseases play out in populations, they said, and can shed light on the movement of agents between farmed and wild fish.
“The tools we employed in this study may prove useful for disease management and fish health. We have shown that for many agents, patterns of infection in dead and dying fish mirror those in live fish,” they explained.
“By integrating high-throughput infectious-agent screening with existing monitoring of dead fish, farm vets and managers could access a wealth of otherwise unavailable or costly information. Combining such results with strategic sacrificial sampling of live fish during mortality peaks could allow additional insight into which agents may be driving mortality.”
To read the full article published in Scientific Reports, click here.






