Genetic sequencing uncovers three new fish virus species
Three new viruses have been discovered in fish as part of genetic sequencing work by PHARMAQ Analytiq, Norwegian University of Life Sciences (NMBU) and University of Minnesota, bringing new knowledge to an emerging field of viral research.
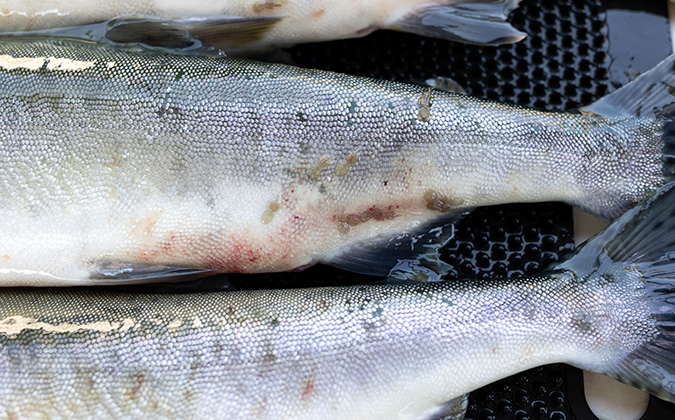
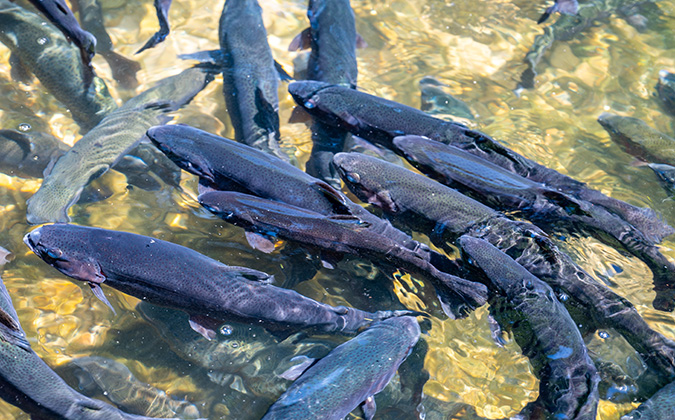
The viruses, known as toti-like viruses due to their similarities to those in the Totiviridae virus family, were detected in farmed lumpsucker in Norway, as well as wild common carp and bluegill from the US.
Comparing the new species to other similar viruses has led the researchers to suggest that the piscine toti-like viruses should form their own genus, Pistolvirus.
“The Totiviridae is known to be a virus family that infects single-celled organisms like fungi and protozoa,” explained Aase Mikalsen, PhD, from NMBU.
“What we call toti-like viruses have been found in invertebrates such as insects, shrimp and crabs, with fish the first vertebrates where this virus has been detected. They have a more complex genome, because they probably need more proteins to be able to infect the more advanced hosts.”
“A common characteristic among the recently discovered toti-like viruses infecting arthropods and fish is that they transmit from the outside of cells and have extra protein-coding sequences that may be connected to their cell entry or exit machineries.”
Lessons for management of salmon pathogen
The newly discovered viruses have not been associated with disease in their fish hosts, but another well-known toti-like virus used as a comparison in the study, piscine myocarditis virus (PMCV), causes serious heart disease in adult Atlantic salmon, with around 100 outbreaks a year in Norway.1
“It’s very significant to the industry to get more knowledge about this virus, because at present there are no vaccines on the market,” Mikalsen said.
“If we can also learn from these other toti-like viruses in fish how they behave and why they are not causing disease in the same way that PMCV does, then you can combine all this knowledge.”
While the viruses detected in the study don’t appear to present the immediate threat to aquaculture of PMCV, Liv Sandlund, PhD, scientist for PHARMAQ Analytiq, sounded a note of caution.
“We don’t know what will happen in the future. We do not know if there are more of these viruses present in the water with the salmon, and whether they can pick up genetic traits that will in the future make the viruses virulent or cause disease,” she said.
Lumpfish importance means surveillance needed
Further studies following up on new viral species, such as Cyclopterus lumpus toti-like virus (CLuTLV) found in lumpfish, are important for fish farming, Mikalsen suggested.
“We cannot just put the knowledge of a virus in a drawer and forget about it. More intensive production might sometimes increase the danger by changing the environment for the virus and fish. It’s important to follow these viruses and the presence and prevalence of them, because they might cause disease later under different conditions,” she continued.
“In particular, the prevalence of CLuTLV is significant, as lumpfish are so widely used in control of sea lice in salmon aquaculture. Though no relationship to disease is clear at present, it’s possible that this could be a matter of virus levels, or that currently only low-virulent variants are circulating in lumpfish production. It is important to continue monitoring this virus, given the increasing production pressure of farmed lumpfish.”
Accessible sequencing technology increases discoveries
The research reflects a broader trend for an increased pace of microbiological discoveries, aided by greater availability of new sequencing technologies.
“We are able to discover more microbes now. Today’s next-generation sequencing platforms are more accessible to both commercial and academic labs and can handle a lot more samples faster and easier. It is also more affordable to do whole-genome sequencing today compared to just a few years ago, and I think that’s why we will continue to see an increase in the discovery of novel viruses and bacterial strains in all fields of research,” Sandlund added.
To view the full study published in Viruses, click here.
1 Fiskehelserapporten 2020; Sommerset I, Bang Jensen B, Bornø G, Haukaas A, Brun E. Eds.; Veterinærinstituttet: Oslo, Norway, 2021.






